Recently, a group of researchers led by Dr. Liandong Qu from Harbin Veterinary Research Institute (HVRI) of Chinese Academy of Agricultural Sciences (CAAS) found a new mechanism by which feline calicivirus (FCV) antagonizes the type IFN response and establishes the persistent infection. This finding suggests that FCV p30 protein can directly degrade IFNAR1 mRNA, which blocks the type I interferon-mediated antiviral response. The article was published in “PLoS Pathogens”.
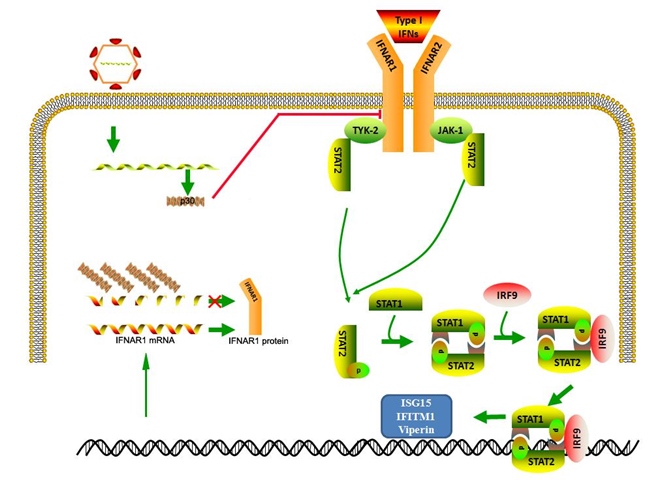
Fig. FCV p30 blocks type I IFN response by directly degrading IFNAR1 mRNA
Most members of the family Caliciviridae can not be cultivated in vitro, and no highly efficient, easy-to-use cell culture model is available. However, FCV (genus Vesivirus) is an excellent model to explore the calicivirus biology due to their culturability and the availability of mature animal models for virus pathogenesis. Once infection has occurred, FCV can persist in the cat population, but the molecular mechanism of how it escapes the innate immune response is still unknown. Here, the authors show that FCV strain 2280 is resistant to the antiviral effect of IFN. The molecular mechanism by which this occurs is that FCV 2280 infection blocks the JAK-STAT pathway through promoting the degradation of IFNAR1 mRNA by FCV p30. An in vitro degradation assay demonstrated that 2280 p30, but not p30 of the vaccine strain F9, could directly and selectively decay IFNAR1 RNA. The exchange of p30 between 2280 and F9 strains using a reverse genetic system also showed that 2280 p30 is a key factor that contributes to the resistance to IFN and enhances virulence. The findings reveal a new mechanism evolved by FCV to circumvent the host antiviral response.
This study was supported by the National Natural Science Foundation of China (grant no.31770172).
More details are available on the link below:
https://journals.plos.org/plospathogens/article?id=10.1371/journal.ppat.1008944.
By Tian Jin(tj6049345@126.com)
|
